※ WocEA INTRODUCTION:
The concept of "Word Clouds" was raised by Jonathan Feinberg, who created the first tool, Wordle, in June 2008 for the visualization of single words or short terms in word wlouds (Viégas et al., 2009). Several years ago, I (Dr. Yu Xue) read an essay written in Chinese by Dr. Wei Li, who works in Netbase Solutions and posted the essay on his ScienceNet blog. Sorry that I forget the contents of the essay, and I cannot find the exact post because Dr. Li posted a lot of blogs, and in several essays he used the Word Cloud layout to show his data, although his posts have no relations with biology. At that time, what I am thinking is whether such a layout can be helpful for visualizing the biological data? So what kinds of biological problems can be resolved by Word Clouds? Perhaps as you know, the direct visualization of individual words or short terms in biology is not frequent, and usually meaningless. I thought this problem for several years.
Now, in my thinkings, I felt one of the most appropriate problems in biology is the visualization of the enrichment analysis, which was raised by Dr. George M. Church, a top-level scientists in a lot of fields, in 2009 (Tavazoie et al., 2009). In functional enrichment analysis, terms or annotations from Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) or other data resources were adopted, and usually the hypergeometric test or similar statistical approaches will be used to test whether a specific term is significantly over-represented (enriched) or under-represented (deprived) in a given list against the background. In 2003, my friend, Dr. Yi Xing, who worked as the Program Director of the Bioinformatics Interdepartmental Graduate Program at the University of California Los Angeles, and now hold the Francis West Lewis Chair in Computational and Genomic Medicine and serve as founder and Director of the Center for Computational and Genomic Medicine at CHOP (See News), published a paper on the impact of alternatively splicing for removing transmembrane domains in protein products (Xing et al., 2003) . He used the hypergeometric distribution in his paper for the enrichment analysis, and I learnt it from him, since I learnt Bioinformatics from him. Two years later, I published a paper to predict sumoylated proteins in human and mouse, and conducted a hypergeometric enrichment analysis for conserved SUMO substrates (Zhou et al., 2005). In early years, the results of enrichment analyses were usually shown in tables, because bioinformaticians did not know how to visualize them at that time.
Thus, here, we developed WocEA (Word cloud for the Enrichment Analysis, version 1.0), mainly for the visualization of enrichment analyses, although the conventional option for showing words or short terms is reserved. Two problems we have resolved are including the proper layout of long terms since GO or KEGG terms are relatively long, and using font size and color to denote the enrichment ratio (E-ratio) and p-value or vice versa, since both two measurements are important output from the hypergeometric test. We anticipate such a layout can visualize the enrichment results in a concise but accurate manner.
Finally, welcome to try it! Please write to us whether you like it or not. We will try our best to address all concerns from you. As all tools and databases of the CUCKOO Workgroup, WocEA will be permenantly free for academic research, and will be continously maintained and updated.
WocEA 1.0 User Interface
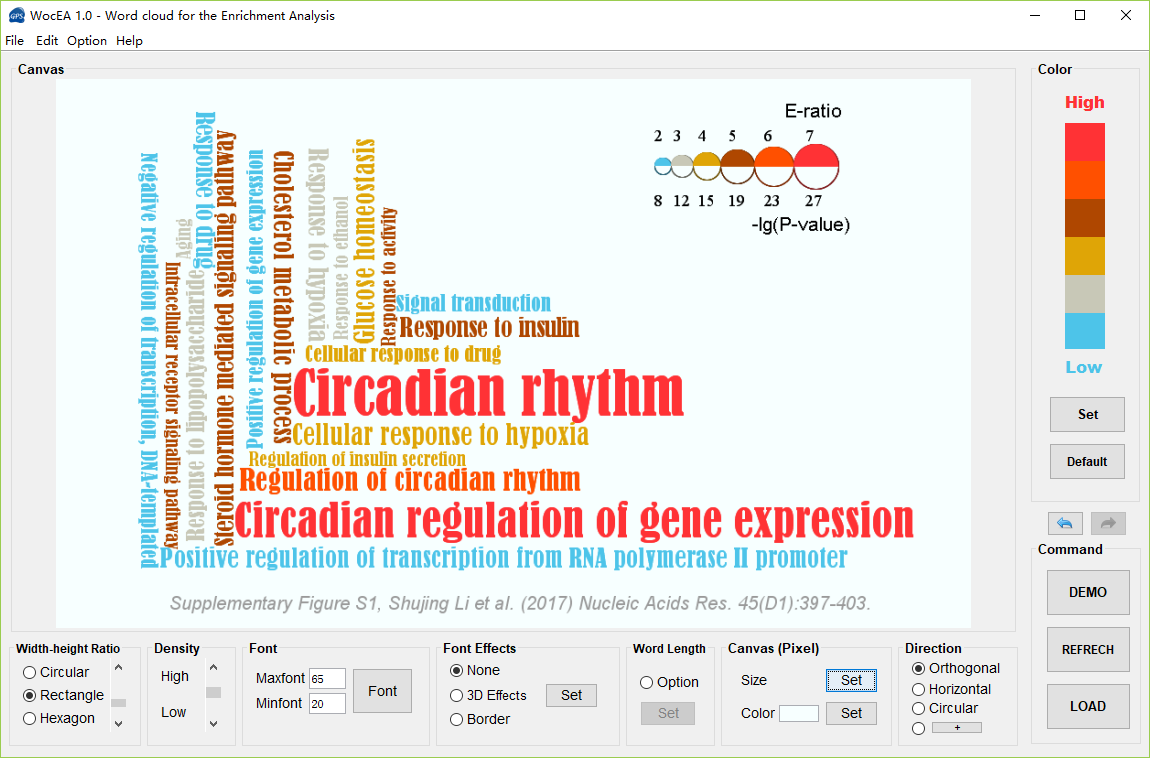
For publication of results please cite the following article:  WocEA: The visualization of functional enrichment results in word clouds
WocEA: The visualization of functional enrichment results in word clouds
Wanshan Ning, Shaofeng Lin, Jiaqi Zhou, Yaping Guo, Ying Zhang, Di Peng and Wankun Deng, Yu Xue. Journal of Genetics and Genomics [Abstract] | [Full Text] | [PDF] |



